```{r setup, include=FALSE}
knitr::opts_chunk$set(echo = TRUE, comment = "", fig.height = 5.5, fig.width = 5.5, cache = TRUE)
```
## Overall Pipeline
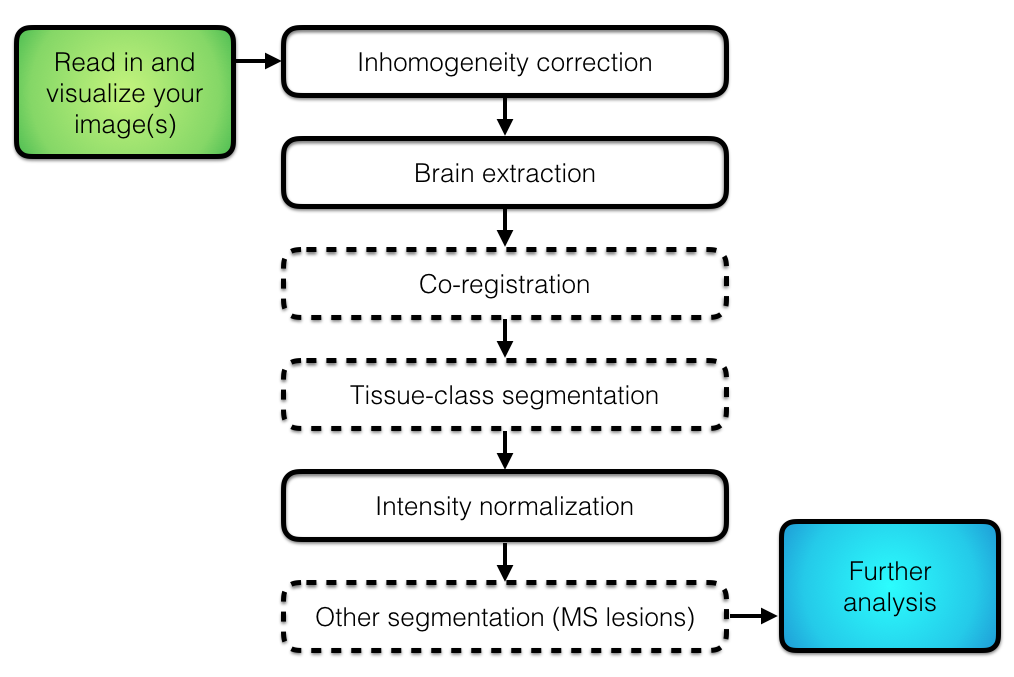 ## Intensity normalization
- Conventional MRI intensites (T1-w, T2-w, PD, FLAIR) are acquired in arbitrary units
- Images are not comparable across scanners, subjects, and visits, even when the same protocol is used.
- This affects algorithm performance, prediction, inference.
- Even simple things like thresholding an image
- Intensity normalization brings the intensities to a common scale across people.
- In this tutorial we will normalize intensities within subject using two methods:
- Whole-brain normalization
- White Stripe normalization [@shinohara2014statistical].
```{r t1, echo=FALSE, warning=FALSE, message=FALSE}
library(ms.lesion)
library(neurobase)
library(WhiteStripe)
fnames = get_image_filenames_list_by_subject(group = "training",
type = "coregistered")
t1s = lapply(fnames, function(x) readnii(x["T1"]))
tissues = lapply(fnames, function(x) readnii(x["Tissue_Classes"]))
masks = lapply(fnames, function(x) readnii(x["Brain_Mask"]))
vals = mapply(function(t1, mask){
mask_vals(t1, mask)
}, t1s, masks, SIMPLIFY = FALSE)
dens = lapply(vals, density)
plot_densities = function(dens, xlab = "Raw Intensities",
main = "Whole Brain") {
range_x = sapply(dens, function(d) range(d$x))
range_x = range(range_x)
range_y = sapply(dens, function(d) range(d$y))
range_y = range(range_y)
plot(dens[[1]], xlim = range_x, ylim = range_y,
xlab = xlab, main = main)
for (idens in 2:length(dens)) {
lines(dens[[idens]], col = idens)
}
}
plot_boxplots = function(vals,
main = "Whole Brain") {
boxplots <- lapply(vals, boxplot, outline = FALSE, plot = FALSE)
boxplots = lapply(boxplots, function(x) x$stats)
boxplots <- do.call(cbind, boxplots)
boxplot(boxplots, main = main)
}
```
## Visualizing whole-brain intensities (each line is a person)
- We will work with the T1-w images from the training data.
- Full brain densities are mixtures of the three tissue class distributions.
```{r t1viz, warning=FALSE, message=FALSE, echo = FALSE}
plot_densities(dens, main = "Distribution of all Voxels in Brain Mask")
```
## Visualizing the intensities by tissue class
```{r threeway, warning=FALSE, message=FALSE, echo = FALSE, fig.height = 4, fig.width = 10}
csf_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 1)
}, t1s, tissues, SIMPLIFY = FALSE)
csf_dens = lapply(csf_vals, density)
wm_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 3)
}, t1s, tissues, SIMPLIFY = FALSE)
wm_dens = lapply(wm_vals, density)
gm_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 2)
}, t1s, tissues, SIMPLIFY = FALSE)
gm_dens = lapply(gm_vals, density)
par(mfrow = c(1, 3))
plot_densities(csf_dens, main = "CSF")
plot_densities(gm_dens, main = "Gray Matter")
plot_densities(wm_dens, main = "White Matter")
par(mfrow = c(1, 1))
```
And these are all the same scanner/protocol!
## Whole-brain normalization
- Let's Z-score each voxel using mean $\mu_{WB}$ and standard deviation $\sigma_{WB}$ computed from all voxels in the brain mask.
$$ T1_{WB} = \frac{T1 - \mu_{WB}}{\sigma_{WB}}$$
- `zscore_img` is a function in `neurobase` that does this.
- It takes an image and a binary mask. The default is to use all voxels in the brain mask.
```{r wbViz, echo=FALSE, warning=FALSE, message=FALSE}
t1_norm = mapply(function(img, mask){
zscore_img(img = img, mask = mask)
}, t1s, masks, SIMPLIFY = FALSE)
```
```{r wbViz_show, eval=FALSE, warning=FALSE, message=FALSE}
zscore_img(img = img, mask = mask)
```
## Whole-brain normalized intensities
## Intensity normalization
- Conventional MRI intensites (T1-w, T2-w, PD, FLAIR) are acquired in arbitrary units
- Images are not comparable across scanners, subjects, and visits, even when the same protocol is used.
- This affects algorithm performance, prediction, inference.
- Even simple things like thresholding an image
- Intensity normalization brings the intensities to a common scale across people.
- In this tutorial we will normalize intensities within subject using two methods:
- Whole-brain normalization
- White Stripe normalization [@shinohara2014statistical].
```{r t1, echo=FALSE, warning=FALSE, message=FALSE}
library(ms.lesion)
library(neurobase)
library(WhiteStripe)
fnames = get_image_filenames_list_by_subject(group = "training",
type = "coregistered")
t1s = lapply(fnames, function(x) readnii(x["T1"]))
tissues = lapply(fnames, function(x) readnii(x["Tissue_Classes"]))
masks = lapply(fnames, function(x) readnii(x["Brain_Mask"]))
vals = mapply(function(t1, mask){
mask_vals(t1, mask)
}, t1s, masks, SIMPLIFY = FALSE)
dens = lapply(vals, density)
plot_densities = function(dens, xlab = "Raw Intensities",
main = "Whole Brain") {
range_x = sapply(dens, function(d) range(d$x))
range_x = range(range_x)
range_y = sapply(dens, function(d) range(d$y))
range_y = range(range_y)
plot(dens[[1]], xlim = range_x, ylim = range_y,
xlab = xlab, main = main)
for (idens in 2:length(dens)) {
lines(dens[[idens]], col = idens)
}
}
plot_boxplots = function(vals,
main = "Whole Brain") {
boxplots <- lapply(vals, boxplot, outline = FALSE, plot = FALSE)
boxplots = lapply(boxplots, function(x) x$stats)
boxplots <- do.call(cbind, boxplots)
boxplot(boxplots, main = main)
}
```
## Visualizing whole-brain intensities (each line is a person)
- We will work with the T1-w images from the training data.
- Full brain densities are mixtures of the three tissue class distributions.
```{r t1viz, warning=FALSE, message=FALSE, echo = FALSE}
plot_densities(dens, main = "Distribution of all Voxels in Brain Mask")
```
## Visualizing the intensities by tissue class
```{r threeway, warning=FALSE, message=FALSE, echo = FALSE, fig.height = 4, fig.width = 10}
csf_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 1)
}, t1s, tissues, SIMPLIFY = FALSE)
csf_dens = lapply(csf_vals, density)
wm_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 3)
}, t1s, tissues, SIMPLIFY = FALSE)
wm_dens = lapply(wm_vals, density)
gm_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 2)
}, t1s, tissues, SIMPLIFY = FALSE)
gm_dens = lapply(gm_vals, density)
par(mfrow = c(1, 3))
plot_densities(csf_dens, main = "CSF")
plot_densities(gm_dens, main = "Gray Matter")
plot_densities(wm_dens, main = "White Matter")
par(mfrow = c(1, 1))
```
And these are all the same scanner/protocol!
## Whole-brain normalization
- Let's Z-score each voxel using mean $\mu_{WB}$ and standard deviation $\sigma_{WB}$ computed from all voxels in the brain mask.
$$ T1_{WB} = \frac{T1 - \mu_{WB}}{\sigma_{WB}}$$
- `zscore_img` is a function in `neurobase` that does this.
- It takes an image and a binary mask. The default is to use all voxels in the brain mask.
```{r wbViz, echo=FALSE, warning=FALSE, message=FALSE}
t1_norm = mapply(function(img, mask){
zscore_img(img = img, mask = mask)
}, t1s, masks, SIMPLIFY = FALSE)
```
```{r wbViz_show, eval=FALSE, warning=FALSE, message=FALSE}
zscore_img(img = img, mask = mask)
```
## Whole-brain normalized intensities
```{r t1viz1a, warning=FALSE, message=FALSE, echo = FALSE}
csf_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 1)
}, t1s, tissues, SIMPLIFY = FALSE)
csf_dens = lapply(csf_vals, density)
plot_densities(csf_dens, main = "CSF Before")
```
```{r wbViz1, warning=FALSE, message=FALSE, echo = FALSE}
csf_norm_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 1)
}, t1_norm, tissues, SIMPLIFY = FALSE)
csf_norm_dens = lapply(csf_norm_vals, density)
plot_densities(csf_norm_dens,
xlab = "Whole-brain Normalized Intensities",
main = "CSF After")
```
## Whole-brain normalized intensities
- Gray matter distributions are more comparable.
```{r t1viz2a, warning=FALSE, message=FALSE, echo = FALSE}
gm_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 2)
}, t1s, tissues, SIMPLIFY = FALSE)
gm_dens = lapply(gm_vals, density)
plot_densities(gm_dens, main = "Gray Matter Before")
```
```{r wbViz2, warning=FALSE, message=FALSE, echo = FALSE}
gm_norm_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 2)
}, t1_norm, tissues, SIMPLIFY = FALSE)
gm_norm_dens = lapply(gm_norm_vals, density)
plot_densities(gm_norm_dens,
xlab = "Whole-brain Normalized Intensities",
main = "Gray Matter After")
```
## Whole-brain normalized intensities
- White matter distributions are more comparable.
```{r t1viz3a, warning=FALSE, message=FALSE, echo = FALSE}
wm_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 3)
}, t1s, tissues, SIMPLIFY = FALSE)
wm_dens = lapply(wm_vals, density)
plot_densities(wm_dens, main = "White Matter Before")
```
```{r wbViz3, warning=FALSE, message=FALSE, echo = FALSE}
wm_norm_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 3)
}, t1_norm, tissues, SIMPLIFY = FALSE)
wm_norm_dens = lapply(wm_norm_vals, density)
plot_densities(wm_norm_dens,
xlab = "Whole-brain Normalized Intensities",
main = "White Matter After")
```
## Other Normalizations: White Stripe
- Whole-brain normalization may be sensitive to outliers.
- Lesions in MS can have very high intensities, which lead to bad estimates of mean/variance
- Other more robust transformations may be used, such as using the median to center, IQR to scale, etc.
- White Stripe [@shinohara2014statistical] is based on parameters obtained from a sample of normal appearing white matter (NAWM), which is robust to outliers.
- The idea is to make normal appearing white matter comparable across subjects and visits.
## White Stripe normalization
Procedure:
1. Find white matter area on histogram
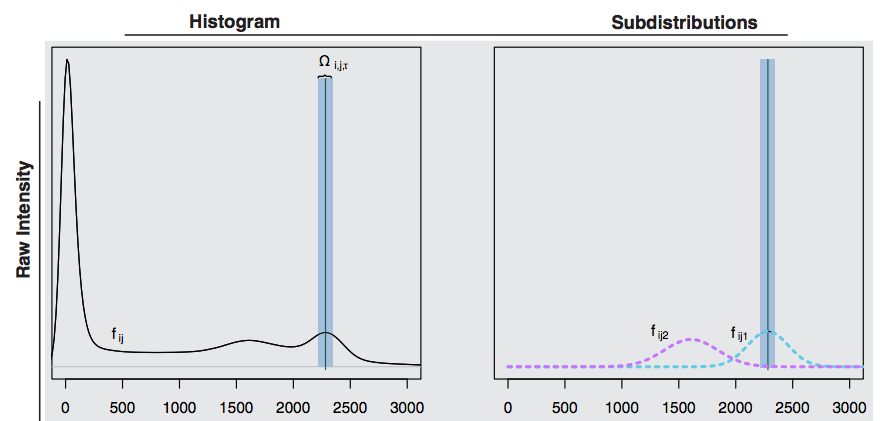 ## White Stripe normalization
Procedure:
1. Find white matter area on histogram
2. Estimate mean $\mu_{WS}$ and variance $\sigma_{WS}$ of voxel intensities in that area
3. Normalize with those means/variances:
$$ T1_{WS} = \frac{T1 - \mu_{WS}}{\sigma_{WS}}$$
## White Stripe normalization
- After normalization, NAWM will have a standard normal distribution and units will be in standard deviations of NAWM.
- Gray matter and CSF distributions may not be comparable after White Stripe.
## White Stripe normalization
Procedure:
1. Find white matter area on histogram
2. Estimate mean $\mu_{WS}$ and variance $\sigma_{WS}$ of voxel intensities in that area
3. Normalize with those means/variances:
$$ T1_{WS} = \frac{T1 - \mu_{WS}}{\sigma_{WS}}$$
## White Stripe normalization
- After normalization, NAWM will have a standard normal distribution and units will be in standard deviations of NAWM.
- Gray matter and CSF distributions may not be comparable after White Stripe.
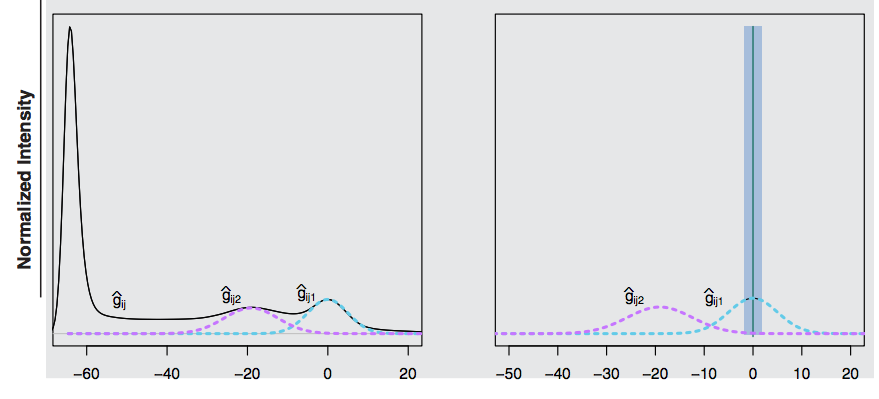 ## White Stripe normalization
Procedure:
1. Find white matter area on histogram
2. Estimate mean $\mu_{WS}$ and variance $\sigma_{WS}$ of voxel intensities in that area
3. Normalize with those means/variances:
$$ T1_{WS} = \frac{T1 - \mu_{WS}}{\sigma_{WS}}$$
- After normalization, NAWM will have a standard normal distribution and units will be in standard deviations of NAWM.
- Gray matter and CSF distributions may not be comparable after White Stripe.
## White Stripe normalization code
```{r ws_show, eval=FALSE, warning = FALSE, message = FALSE, results='hide'}
library(WhiteStripe)
ind = whitestripe(img = t1, type = "T1", stripped = TRUE)$whitestripe.ind
ws_t1 = whitestripe_norm(t1, indices = ind)
```
- The `whitestripe` function takes an image, image type (in our case T1), and a logical indicating whether the image has been skull stripped.
- The indicies of voxels in the NAWM used for estimating the normalization parameters are located in the list element `$whitestripe.ind`.
- The function `whitestripe_norm` takes an image and the indicies from a call to `whitestripe` and returns the White Stripe normalized image as a nifti.
```{r ws, echo=FALSE, warning = FALSE, message = FALSE, results='hide'}
ws_norm = function(t1) {
ind = whitestripe(img = t1,
type = "T1",
stripped = TRUE)$whitestripe.ind
whitestripe_norm(t1, indices = ind)
}
t1_ws_norm = lapply(t1s, ws_norm)
```
## WhiteStripe normalized intensities
## White Stripe normalization
Procedure:
1. Find white matter area on histogram
2. Estimate mean $\mu_{WS}$ and variance $\sigma_{WS}$ of voxel intensities in that area
3. Normalize with those means/variances:
$$ T1_{WS} = \frac{T1 - \mu_{WS}}{\sigma_{WS}}$$
- After normalization, NAWM will have a standard normal distribution and units will be in standard deviations of NAWM.
- Gray matter and CSF distributions may not be comparable after White Stripe.
## White Stripe normalization code
```{r ws_show, eval=FALSE, warning = FALSE, message = FALSE, results='hide'}
library(WhiteStripe)
ind = whitestripe(img = t1, type = "T1", stripped = TRUE)$whitestripe.ind
ws_t1 = whitestripe_norm(t1, indices = ind)
```
- The `whitestripe` function takes an image, image type (in our case T1), and a logical indicating whether the image has been skull stripped.
- The indicies of voxels in the NAWM used for estimating the normalization parameters are located in the list element `$whitestripe.ind`.
- The function `whitestripe_norm` takes an image and the indicies from a call to `whitestripe` and returns the White Stripe normalized image as a nifti.
```{r ws, echo=FALSE, warning = FALSE, message = FALSE, results='hide'}
ws_norm = function(t1) {
ind = whitestripe(img = t1,
type = "T1",
stripped = TRUE)$whitestripe.ind
whitestripe_norm(t1, indices = ind)
}
t1_ws_norm = lapply(t1s, ws_norm)
```
## WhiteStripe normalized intensities
```{r wbViz1a, warning=FALSE, message=FALSE, echo = FALSE}
plot_densities(csf_norm_dens,
xlab = "Whole-brain Normalized Intensities", main = "Whole-brain: CSF")
```
```{r ws_viz_csf, warning=FALSE, message=FALSE, echo = FALSE}
csf_ws_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 1)
}, t1_ws_norm, tissues, SIMPLIFY = FALSE)
csf_ws_dens = lapply(csf_ws_vals, density)
plot_densities(csf_ws_dens,
xlab = "WhiteStripe Normalized Intensities", main = "White Stripe: CSF")
```
## WhiteStripe normalized intensities
```{r wbViz2a, warning=FALSE, message=FALSE, echo = FALSE}
plot_densities(gm_norm_dens,
xlab = "Whole-brain Normalized Intensities", main = "Whole-brain: Gray Matter")
```
```{r ws_viz_gm, warning=FALSE, message=FALSE, echo = FALSE}
gm_ws_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 2)
}, t1_ws_norm, tissues, SIMPLIFY = FALSE)
gm_ws_dens = lapply(gm_ws_vals, density)
plot_densities(gm_ws_dens,
xlab = "WhiteStripe Normalized Intensities", main = "White Stripe: Gray Matter")
```
## WhiteStripe normalized intensities
```{r wbViz3a, warning=FALSE, message=FALSE, echo = FALSE}
plot_densities(wm_norm_dens,
xlab = "Whole-brain Normalized Intensities", main = "Whole-brain: White Matter")
```
```{r ws_viz_wm, warning=FALSE, message=FALSE, echo = FALSE}
wm_ws_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 3)
}, t1_ws_norm, tissues, SIMPLIFY = FALSE)
wm_ws_dens = lapply(wm_ws_vals, density)
plot_densities(wm_ws_dens,
xlab = "WhiteStripe Normalized Intensities", main = "White Stripe: White Matter")
```
## Conclusions
- Intensity normalization is an important step in any image analysis with more than one subject or time point to ensure comparability across images.
- White Stripe normalization may work better and have better interpretation than whole-brain normalization for subsequent lesion segmentation algorithms and analysis.
- Other intensity normalization methods that make intensites comparable across subjects for all tissues exist.
- RAVEL, which is an extension of WhiteStripe is one example [@fortin2016removing].
- Located at https://github.com/Jfortin1/RAVEL
- This was shown to have better comparability than histogram matching
## Website
http://johnmuschelli.com/imaging_in_r
## References {.smaller}
 ## Intensity normalization
- Conventional MRI intensites (T1-w, T2-w, PD, FLAIR) are acquired in arbitrary units
- Images are not comparable across scanners, subjects, and visits, even when the same protocol is used.
- This affects algorithm performance, prediction, inference.
- Even simple things like thresholding an image
- Intensity normalization brings the intensities to a common scale across people.
- In this tutorial we will normalize intensities within subject using two methods:
- Whole-brain normalization
- White Stripe normalization [@shinohara2014statistical].
```{r t1, echo=FALSE, warning=FALSE, message=FALSE}
library(ms.lesion)
library(neurobase)
library(WhiteStripe)
fnames = get_image_filenames_list_by_subject(group = "training",
type = "coregistered")
t1s = lapply(fnames, function(x) readnii(x["T1"]))
tissues = lapply(fnames, function(x) readnii(x["Tissue_Classes"]))
masks = lapply(fnames, function(x) readnii(x["Brain_Mask"]))
vals = mapply(function(t1, mask){
mask_vals(t1, mask)
}, t1s, masks, SIMPLIFY = FALSE)
dens = lapply(vals, density)
plot_densities = function(dens, xlab = "Raw Intensities",
main = "Whole Brain") {
range_x = sapply(dens, function(d) range(d$x))
range_x = range(range_x)
range_y = sapply(dens, function(d) range(d$y))
range_y = range(range_y)
plot(dens[[1]], xlim = range_x, ylim = range_y,
xlab = xlab, main = main)
for (idens in 2:length(dens)) {
lines(dens[[idens]], col = idens)
}
}
plot_boxplots = function(vals,
main = "Whole Brain") {
boxplots <- lapply(vals, boxplot, outline = FALSE, plot = FALSE)
boxplots = lapply(boxplots, function(x) x$stats)
boxplots <- do.call(cbind, boxplots)
boxplot(boxplots, main = main)
}
```
## Visualizing whole-brain intensities (each line is a person)
- We will work with the T1-w images from the training data.
- Full brain densities are mixtures of the three tissue class distributions.
```{r t1viz, warning=FALSE, message=FALSE, echo = FALSE}
plot_densities(dens, main = "Distribution of all Voxels in Brain Mask")
```
## Visualizing the intensities by tissue class
```{r threeway, warning=FALSE, message=FALSE, echo = FALSE, fig.height = 4, fig.width = 10}
csf_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 1)
}, t1s, tissues, SIMPLIFY = FALSE)
csf_dens = lapply(csf_vals, density)
wm_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 3)
}, t1s, tissues, SIMPLIFY = FALSE)
wm_dens = lapply(wm_vals, density)
gm_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 2)
}, t1s, tissues, SIMPLIFY = FALSE)
gm_dens = lapply(gm_vals, density)
par(mfrow = c(1, 3))
plot_densities(csf_dens, main = "CSF")
plot_densities(gm_dens, main = "Gray Matter")
plot_densities(wm_dens, main = "White Matter")
par(mfrow = c(1, 1))
```
And these are all the same scanner/protocol!
## Whole-brain normalization
- Let's Z-score each voxel using mean $\mu_{WB}$ and standard deviation $\sigma_{WB}$ computed from all voxels in the brain mask.
$$ T1_{WB} = \frac{T1 - \mu_{WB}}{\sigma_{WB}}$$
- `zscore_img` is a function in `neurobase` that does this.
- It takes an image and a binary mask. The default is to use all voxels in the brain mask.
```{r wbViz, echo=FALSE, warning=FALSE, message=FALSE}
t1_norm = mapply(function(img, mask){
zscore_img(img = img, mask = mask)
}, t1s, masks, SIMPLIFY = FALSE)
```
```{r wbViz_show, eval=FALSE, warning=FALSE, message=FALSE}
zscore_img(img = img, mask = mask)
```
## Whole-brain normalized intensities
## Intensity normalization
- Conventional MRI intensites (T1-w, T2-w, PD, FLAIR) are acquired in arbitrary units
- Images are not comparable across scanners, subjects, and visits, even when the same protocol is used.
- This affects algorithm performance, prediction, inference.
- Even simple things like thresholding an image
- Intensity normalization brings the intensities to a common scale across people.
- In this tutorial we will normalize intensities within subject using two methods:
- Whole-brain normalization
- White Stripe normalization [@shinohara2014statistical].
```{r t1, echo=FALSE, warning=FALSE, message=FALSE}
library(ms.lesion)
library(neurobase)
library(WhiteStripe)
fnames = get_image_filenames_list_by_subject(group = "training",
type = "coregistered")
t1s = lapply(fnames, function(x) readnii(x["T1"]))
tissues = lapply(fnames, function(x) readnii(x["Tissue_Classes"]))
masks = lapply(fnames, function(x) readnii(x["Brain_Mask"]))
vals = mapply(function(t1, mask){
mask_vals(t1, mask)
}, t1s, masks, SIMPLIFY = FALSE)
dens = lapply(vals, density)
plot_densities = function(dens, xlab = "Raw Intensities",
main = "Whole Brain") {
range_x = sapply(dens, function(d) range(d$x))
range_x = range(range_x)
range_y = sapply(dens, function(d) range(d$y))
range_y = range(range_y)
plot(dens[[1]], xlim = range_x, ylim = range_y,
xlab = xlab, main = main)
for (idens in 2:length(dens)) {
lines(dens[[idens]], col = idens)
}
}
plot_boxplots = function(vals,
main = "Whole Brain") {
boxplots <- lapply(vals, boxplot, outline = FALSE, plot = FALSE)
boxplots = lapply(boxplots, function(x) x$stats)
boxplots <- do.call(cbind, boxplots)
boxplot(boxplots, main = main)
}
```
## Visualizing whole-brain intensities (each line is a person)
- We will work with the T1-w images from the training data.
- Full brain densities are mixtures of the three tissue class distributions.
```{r t1viz, warning=FALSE, message=FALSE, echo = FALSE}
plot_densities(dens, main = "Distribution of all Voxels in Brain Mask")
```
## Visualizing the intensities by tissue class
```{r threeway, warning=FALSE, message=FALSE, echo = FALSE, fig.height = 4, fig.width = 10}
csf_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 1)
}, t1s, tissues, SIMPLIFY = FALSE)
csf_dens = lapply(csf_vals, density)
wm_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 3)
}, t1s, tissues, SIMPLIFY = FALSE)
wm_dens = lapply(wm_vals, density)
gm_vals = mapply(function(t1, mask){
mask_vals(t1, mask == 2)
}, t1s, tissues, SIMPLIFY = FALSE)
gm_dens = lapply(gm_vals, density)
par(mfrow = c(1, 3))
plot_densities(csf_dens, main = "CSF")
plot_densities(gm_dens, main = "Gray Matter")
plot_densities(wm_dens, main = "White Matter")
par(mfrow = c(1, 1))
```
And these are all the same scanner/protocol!
## Whole-brain normalization
- Let's Z-score each voxel using mean $\mu_{WB}$ and standard deviation $\sigma_{WB}$ computed from all voxels in the brain mask.
$$ T1_{WB} = \frac{T1 - \mu_{WB}}{\sigma_{WB}}$$
- `zscore_img` is a function in `neurobase` that does this.
- It takes an image and a binary mask. The default is to use all voxels in the brain mask.
```{r wbViz, echo=FALSE, warning=FALSE, message=FALSE}
t1_norm = mapply(function(img, mask){
zscore_img(img = img, mask = mask)
}, t1s, masks, SIMPLIFY = FALSE)
```
```{r wbViz_show, eval=FALSE, warning=FALSE, message=FALSE}
zscore_img(img = img, mask = mask)
```
## Whole-brain normalized intensities
 ## White Stripe normalization
Procedure:
1. Find white matter area on histogram
2. Estimate mean $\mu_{WS}$ and variance $\sigma_{WS}$ of voxel intensities in that area
3. Normalize with those means/variances:
$$ T1_{WS} = \frac{T1 - \mu_{WS}}{\sigma_{WS}}$$
## White Stripe normalization
- After normalization, NAWM will have a standard normal distribution and units will be in standard deviations of NAWM.
- Gray matter and CSF distributions may not be comparable after White Stripe.
## White Stripe normalization
Procedure:
1. Find white matter area on histogram
2. Estimate mean $\mu_{WS}$ and variance $\sigma_{WS}$ of voxel intensities in that area
3. Normalize with those means/variances:
$$ T1_{WS} = \frac{T1 - \mu_{WS}}{\sigma_{WS}}$$
## White Stripe normalization
- After normalization, NAWM will have a standard normal distribution and units will be in standard deviations of NAWM.
- Gray matter and CSF distributions may not be comparable after White Stripe.
 ## White Stripe normalization
Procedure:
1. Find white matter area on histogram
2. Estimate mean $\mu_{WS}$ and variance $\sigma_{WS}$ of voxel intensities in that area
3. Normalize with those means/variances:
$$ T1_{WS} = \frac{T1 - \mu_{WS}}{\sigma_{WS}}$$
- After normalization, NAWM will have a standard normal distribution and units will be in standard deviations of NAWM.
- Gray matter and CSF distributions may not be comparable after White Stripe.
## White Stripe normalization code
```{r ws_show, eval=FALSE, warning = FALSE, message = FALSE, results='hide'}
library(WhiteStripe)
ind = whitestripe(img = t1, type = "T1", stripped = TRUE)$whitestripe.ind
ws_t1 = whitestripe_norm(t1, indices = ind)
```
- The `whitestripe` function takes an image, image type (in our case T1), and a logical indicating whether the image has been skull stripped.
- The indicies of voxels in the NAWM used for estimating the normalization parameters are located in the list element `$whitestripe.ind`.
- The function `whitestripe_norm` takes an image and the indicies from a call to `whitestripe` and returns the White Stripe normalized image as a nifti.
```{r ws, echo=FALSE, warning = FALSE, message = FALSE, results='hide'}
ws_norm = function(t1) {
ind = whitestripe(img = t1,
type = "T1",
stripped = TRUE)$whitestripe.ind
whitestripe_norm(t1, indices = ind)
}
t1_ws_norm = lapply(t1s, ws_norm)
```
## WhiteStripe normalized intensities
## White Stripe normalization
Procedure:
1. Find white matter area on histogram
2. Estimate mean $\mu_{WS}$ and variance $\sigma_{WS}$ of voxel intensities in that area
3. Normalize with those means/variances:
$$ T1_{WS} = \frac{T1 - \mu_{WS}}{\sigma_{WS}}$$
- After normalization, NAWM will have a standard normal distribution and units will be in standard deviations of NAWM.
- Gray matter and CSF distributions may not be comparable after White Stripe.
## White Stripe normalization code
```{r ws_show, eval=FALSE, warning = FALSE, message = FALSE, results='hide'}
library(WhiteStripe)
ind = whitestripe(img = t1, type = "T1", stripped = TRUE)$whitestripe.ind
ws_t1 = whitestripe_norm(t1, indices = ind)
```
- The `whitestripe` function takes an image, image type (in our case T1), and a logical indicating whether the image has been skull stripped.
- The indicies of voxels in the NAWM used for estimating the normalization parameters are located in the list element `$whitestripe.ind`.
- The function `whitestripe_norm` takes an image and the indicies from a call to `whitestripe` and returns the White Stripe normalized image as a nifti.
```{r ws, echo=FALSE, warning = FALSE, message = FALSE, results='hide'}
ws_norm = function(t1) {
ind = whitestripe(img = t1,
type = "T1",
stripped = TRUE)$whitestripe.ind
whitestripe_norm(t1, indices = ind)
}
t1_ws_norm = lapply(t1s, ws_norm)
```
## WhiteStripe normalized intensities