```{r setup, include=FALSE}
library(methods)
library(neurobase)
library(extrantsr)
knitr::opts_chunk$set(echo = TRUE, comment = "")
```
## Overall Pipeline
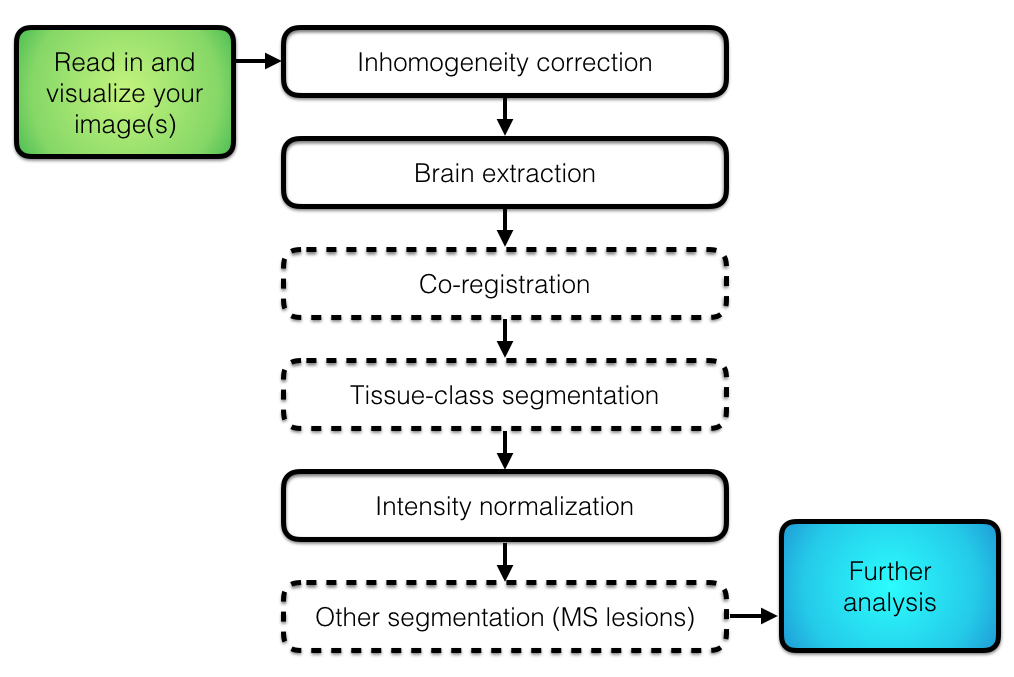 ## Types of Registration
- Rigid-body registration (linear) - 6 degrees of freedom (dof)
## Types of Registration
- Rigid-body registration (linear) - 6 degrees of freedom (dof)
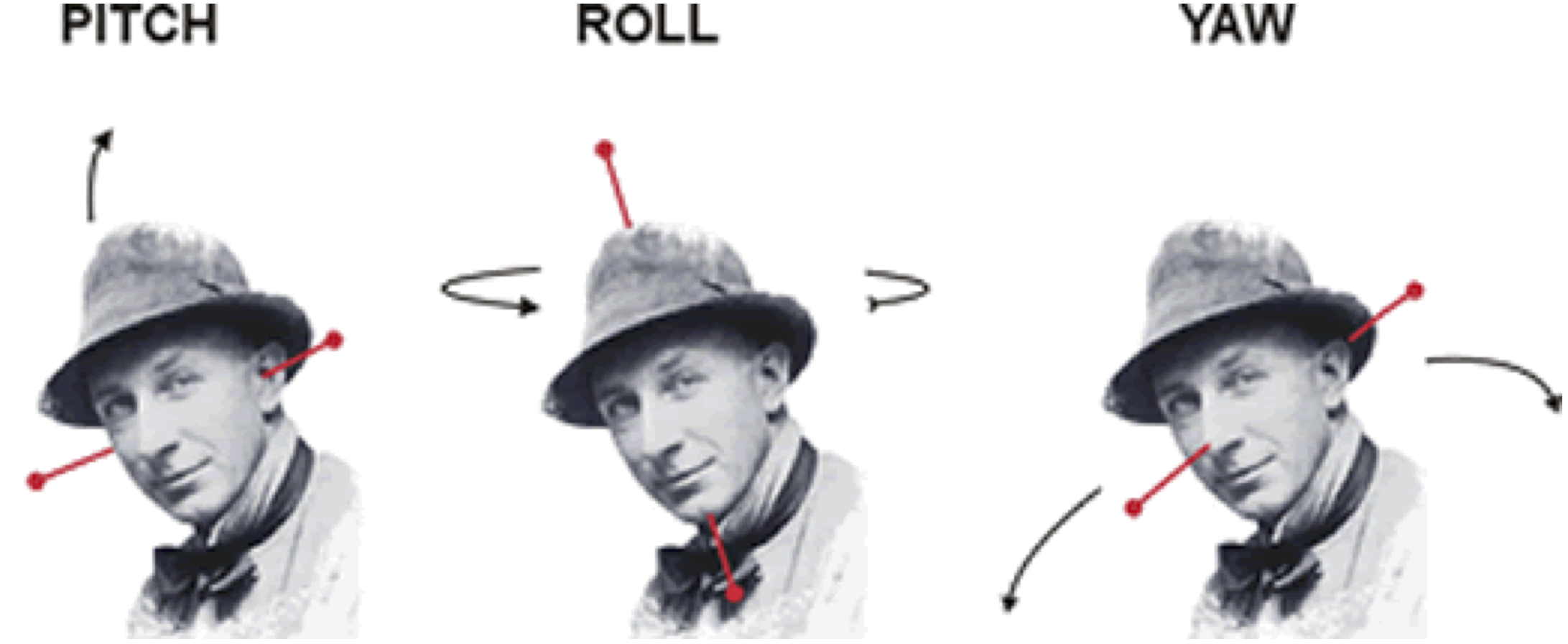
Image taken from [http://cnl.web.arizona.edu/imageprops.htm](http://cnl.web.arizona.edu/imageprops.htm)
- Pitch - Think of nodding ("yes")
- Yaw - Think of shaking head ("no")
- Roll - Think of shoulder shrugging ("I don't know")
- x – left/right
- y – forward/backward
- z – jump up/down
## Types of Registration
- Rigid-body registration (linear) - 6 degrees of freedom (dof)
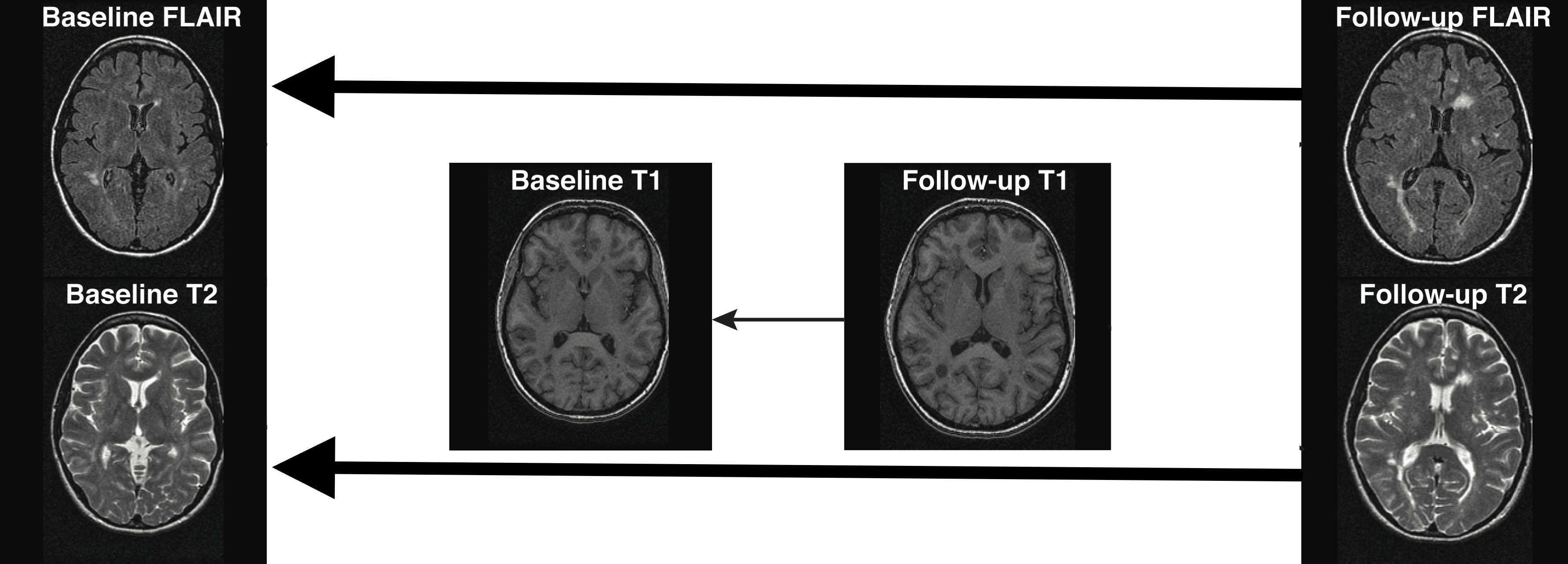
- Co-registration (within the same person)
- **Cross-sectional between-sequences**
- Longitudinal between-sequences
- Longitudinal within-sequence
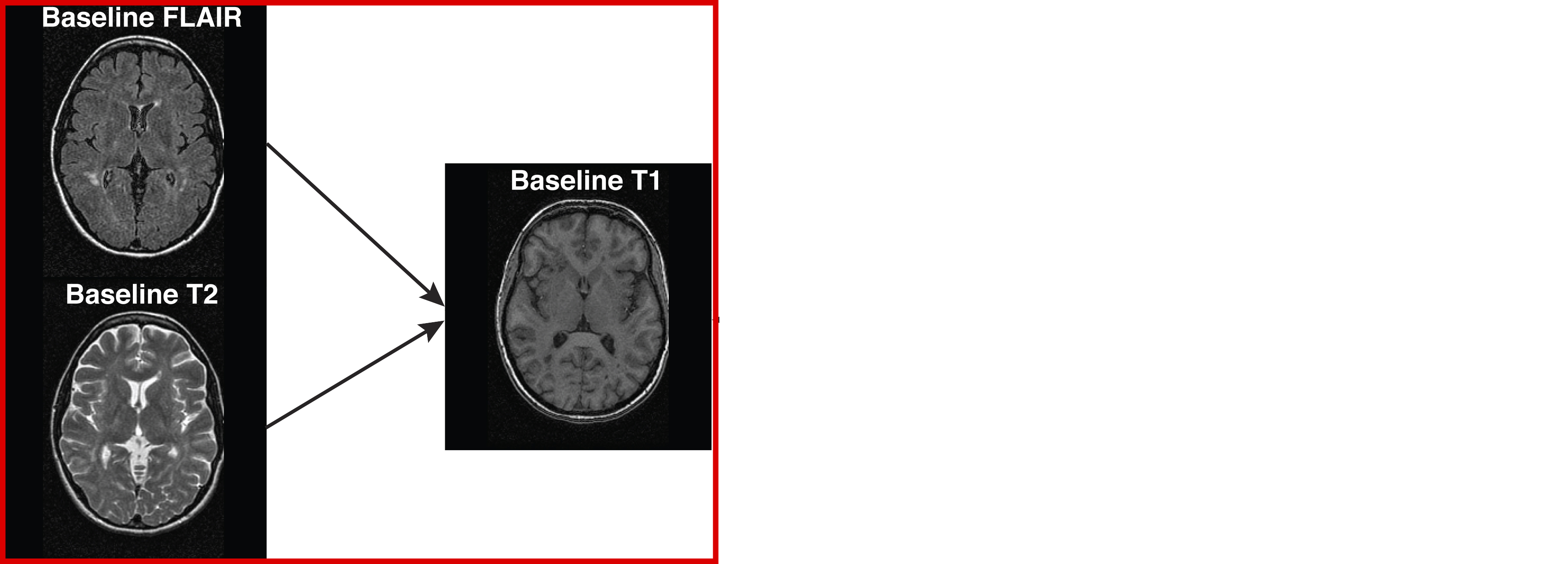 ## Types of Registration
- Rigid-body registration (linear) - 6 degrees of freedom (dof)
- Co-registration (within the same person)
- **Cross-sectional between-sequences**
- **Longitudinal between-sequences**
- Longitudinal within-sequence
## Types of Registration
- Rigid-body registration (linear) - 6 degrees of freedom (dof)
- Co-registration (within the same person)
- **Cross-sectional between-sequences**
- **Longitudinal between-sequences**
- Longitudinal within-sequence
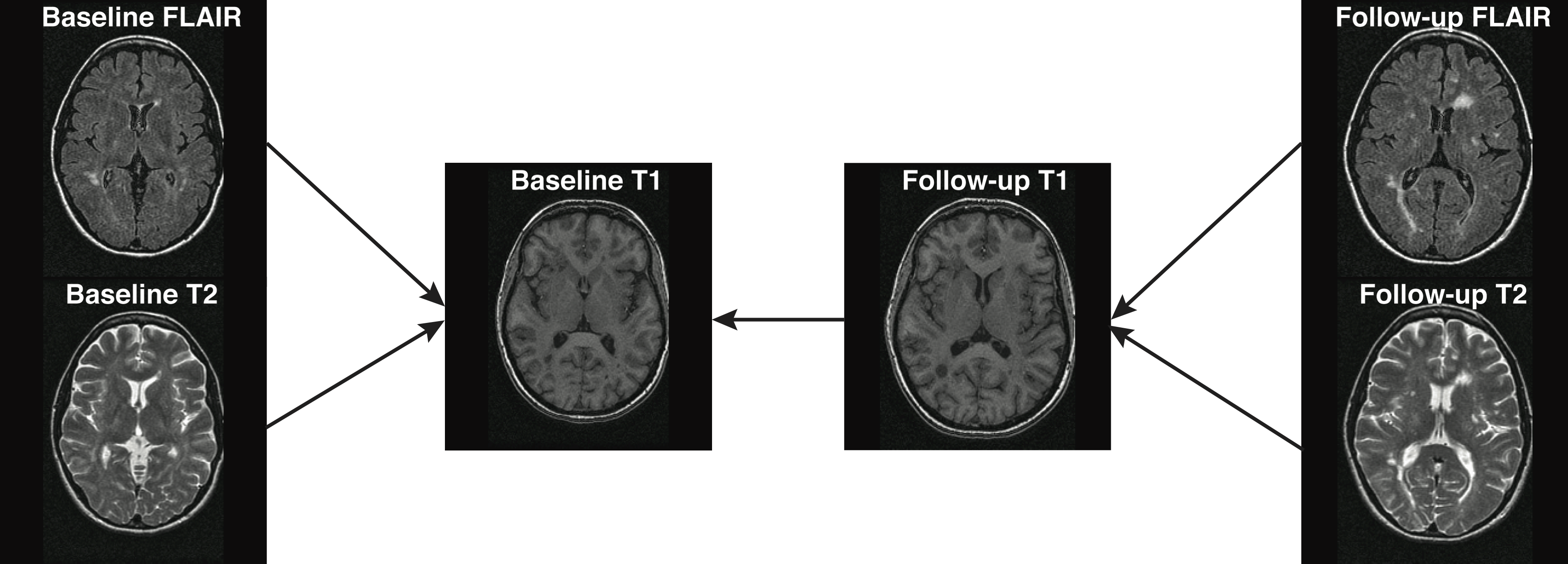 ## Types of Registration
- Rigid-body registration (linear) - 6 degrees of freedom (dof)
- Co-registration (within the same person)
- Cross-sectional between-sequences**
- Longitudinal between-sequences
- **Longitudinal within-sequence**
## Types of Registration
- Rigid-body registration (linear) - 6 degrees of freedom (dof)
- Co-registration (within the same person)
- Cross-sectional between-sequences**
- Longitudinal between-sequences
- **Longitudinal within-sequence**
 ## Types of Registration
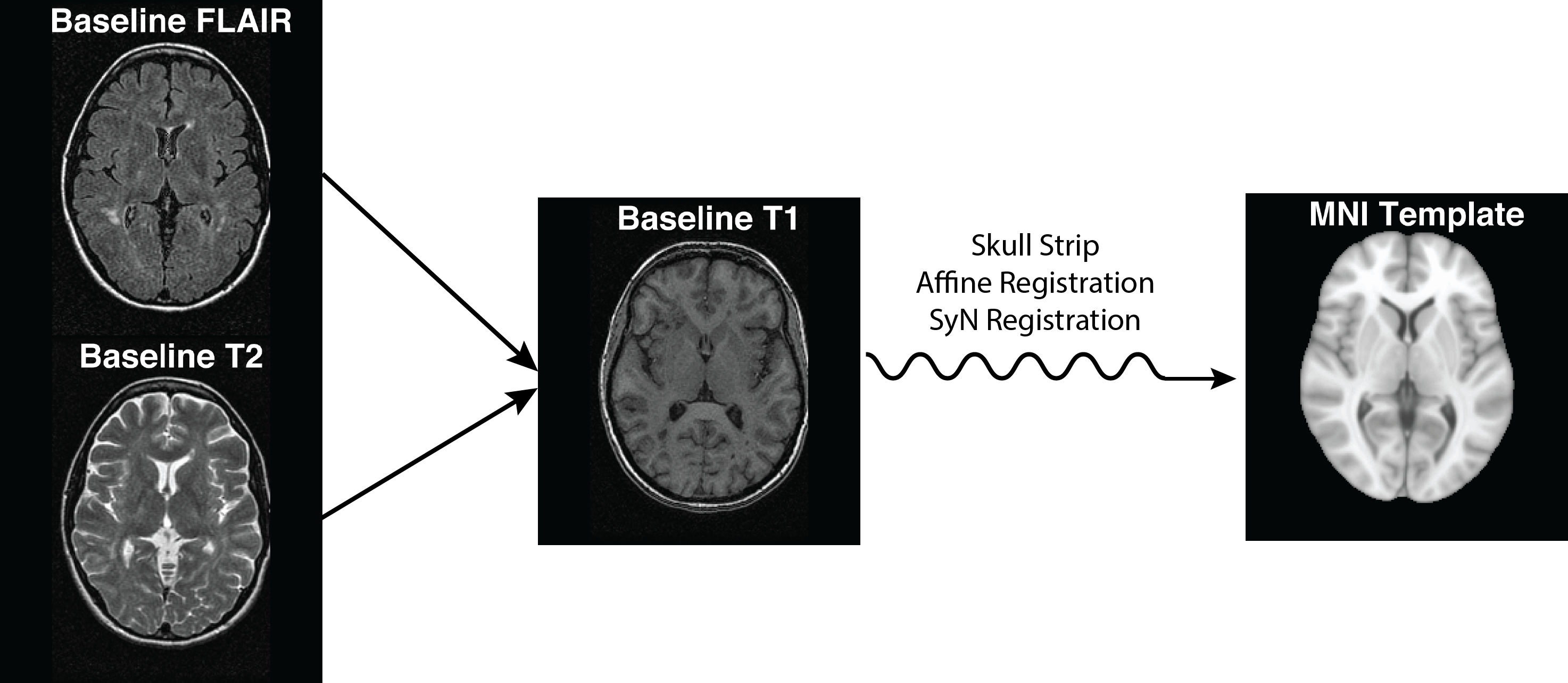
- Affine registration – 12 dof (scaling)
- Non-linear (> 12 dof): sually requires a prior affine registration
- Across-subject registration
- Registration to a template: There are many different templates
## Types of Registration
- Affine registration – 12 dof (scaling)
- Non-linear (> 12 dof): sually requires a prior affine registration
- Across-subject registration
- Registration to a template: There are many different templates
 ## Rigid Registration: The Math
For a voxel $v$, the rigid transformation can be written as:
$$T_{\rm rigid}(v) = Rv + t$$
where $R =$
\small
$$\left[\begin{array}{ccc} \cos\beta\cos\gamma& \cos\alpha\sin\gamma + \sin\alpha\sin\beta\cos\gamma & \sin\alpha\sin\gamma - \cos\alpha\sin\beta\cos\gamma \\
-\cos\beta\sin\gamma & \cos\alpha\cos\gamma - \sin\alpha\sin\beta\sin\gamma & \sin\alpha\cos\gamma + \cos\alpha\sin\beta\sin\gamma \\
\sin\beta & -\sin\alpha\cos\beta & \cos\alpha\cos\beta \end{array}\right]$$
\normalsize
- 6 degrees of freedom
- $3$ associated with the translation vector: $t=(t_x, t_y, t_z)$
- $3$ associated with the rotation parameters: $\theta=(\alpha, \beta,\gamma)$.
## Aside into Lists
- Initialize an empty list and add two elements to it
```{r list}
l = list()
l[[1]] = c(1, 2, 4, 5)
l[[2]] = matrix(1:10, nrow = 2)
print(l)
```
- Subsetting uses double brackets:
```{r listSub}
print(l[[1]])
```
## Subsetting by name
If a `vector` has names, you can also put the
- Initialize an empty list and add two elements to it
```{r named_vec}
x = c(one = 1, three = 14, two = 5)
print(x)
x[c("three")]
```
If a `list` has names, you can subset with the `$`
- Subsetting with double brackets:
```{r named_lst}
names(l) = c("V", "m"); l[["V"]]
```
## Subsetting by name
If a `list` has names, you can also subset with the `$`
```{r named_lst_v}
l$V
```
## Reading in more than the T1: `ms.lesion` package
- `get_image_filenames_list_by_subject` - returns a list, each element is a subject, and each subject has a vector of filenames, named for each modality
```{r t1, message=FALSE}
library(ms.lesion)
all_files = get_image_filenames_list_by_subject()
class(all_files); names(all_files)
files = all_files$training01; class(files); names(files)
t1_fname = files["T1"]
t1 = readnii(t1_fname)
rt1 = robust_window(t1, probs = c(0, 0.975))
```
## Image Registration
The `registration` function from `extrantsr` can register 2 images. The main arguments are:
- `filename` - either `nifti` object or filename of image to be registered (moving)
- `template.file` - either `nifti` object or filename of target image (fixed)
- `typeofTransform` - transformation of moving to fixed image (Rigid/Affine/SyN)
- `interpolator` - how are voxels averaged in `fixed` space (Linear/NearestNeighbor/LanczosWindowedSinc)
It can also perform bias correction if `correct = TRUE`.
## Image registration
For example, if we wanted to register the FLAIR to the T1 image, we would run:
```{r, eval = TRUE, cache = TRUE, message=FALSE}
library(extrantsr)
reg = registration(
filename = files["FLAIR"], template.file = files["T1"],
typeofTransform = "Rigid", interpolator = "Linear", verbose = FALSE)
names(reg)
```
The output in `reg` would contain the transformed image and paths to the estimated transformations.
## FLAIR comparison
```{r plot_reg, eval = FALSE, cache = FALSE, message=FALSE}
double_ortho(rt1, reg$outfile)
```
```{r plot_reg_show, echo = FALSE, eval = TRUE, cache = TRUE, message=FALSE}
reg$outfile = robust_window(reg$outfile)
double_ortho(rt1, reg$outfile)
```
## Wrapper function to perform preprocessing
We would like to perform registration within a visit for all modalities. The `extrantsr` function `within_visit_registration` will do the following steps:
1. Inhomogeneity correction (N3 or N4): could also use images from previous lecture
2. Registration to a fixed image (T1)
## Rigid Registration: The Math
For a voxel $v$, the rigid transformation can be written as:
$$T_{\rm rigid}(v) = Rv + t$$
where $R =$
\small
$$\left[\begin{array}{ccc} \cos\beta\cos\gamma& \cos\alpha\sin\gamma + \sin\alpha\sin\beta\cos\gamma & \sin\alpha\sin\gamma - \cos\alpha\sin\beta\cos\gamma \\
-\cos\beta\sin\gamma & \cos\alpha\cos\gamma - \sin\alpha\sin\beta\sin\gamma & \sin\alpha\cos\gamma + \cos\alpha\sin\beta\sin\gamma \\
\sin\beta & -\sin\alpha\cos\beta & \cos\alpha\cos\beta \end{array}\right]$$
\normalsize
- 6 degrees of freedom
- $3$ associated with the translation vector: $t=(t_x, t_y, t_z)$
- $3$ associated with the rotation parameters: $\theta=(\alpha, \beta,\gamma)$.
## Aside into Lists
- Initialize an empty list and add two elements to it
```{r list}
l = list()
l[[1]] = c(1, 2, 4, 5)
l[[2]] = matrix(1:10, nrow = 2)
print(l)
```
- Subsetting uses double brackets:
```{r listSub}
print(l[[1]])
```
## Subsetting by name
If a `vector` has names, you can also put the
- Initialize an empty list and add two elements to it
```{r named_vec}
x = c(one = 1, three = 14, two = 5)
print(x)
x[c("three")]
```
If a `list` has names, you can subset with the `$`
- Subsetting with double brackets:
```{r named_lst}
names(l) = c("V", "m"); l[["V"]]
```
## Subsetting by name
If a `list` has names, you can also subset with the `$`
```{r named_lst_v}
l$V
```
## Reading in more than the T1: `ms.lesion` package
- `get_image_filenames_list_by_subject` - returns a list, each element is a subject, and each subject has a vector of filenames, named for each modality
```{r t1, message=FALSE}
library(ms.lesion)
all_files = get_image_filenames_list_by_subject()
class(all_files); names(all_files)
files = all_files$training01; class(files); names(files)
t1_fname = files["T1"]
t1 = readnii(t1_fname)
rt1 = robust_window(t1, probs = c(0, 0.975))
```
## Image Registration
The `registration` function from `extrantsr` can register 2 images. The main arguments are:
- `filename` - either `nifti` object or filename of image to be registered (moving)
- `template.file` - either `nifti` object or filename of target image (fixed)
- `typeofTransform` - transformation of moving to fixed image (Rigid/Affine/SyN)
- `interpolator` - how are voxels averaged in `fixed` space (Linear/NearestNeighbor/LanczosWindowedSinc)
It can also perform bias correction if `correct = TRUE`.
## Image registration
For example, if we wanted to register the FLAIR to the T1 image, we would run:
```{r, eval = TRUE, cache = TRUE, message=FALSE}
library(extrantsr)
reg = registration(
filename = files["FLAIR"], template.file = files["T1"],
typeofTransform = "Rigid", interpolator = "Linear", verbose = FALSE)
names(reg)
```
The output in `reg` would contain the transformed image and paths to the estimated transformations.
## FLAIR comparison
```{r plot_reg, eval = FALSE, cache = FALSE, message=FALSE}
double_ortho(rt1, reg$outfile)
```
```{r plot_reg_show, echo = FALSE, eval = TRUE, cache = TRUE, message=FALSE}
reg$outfile = robust_window(reg$outfile)
double_ortho(rt1, reg$outfile)
```
## Wrapper function to perform preprocessing
We would like to perform registration within a visit for all modalities. The `extrantsr` function `within_visit_registration` will do the following steps:
1. Inhomogeneity correction (N3 or N4): could also use images from previous lecture
2. Registration to a fixed image (T1)
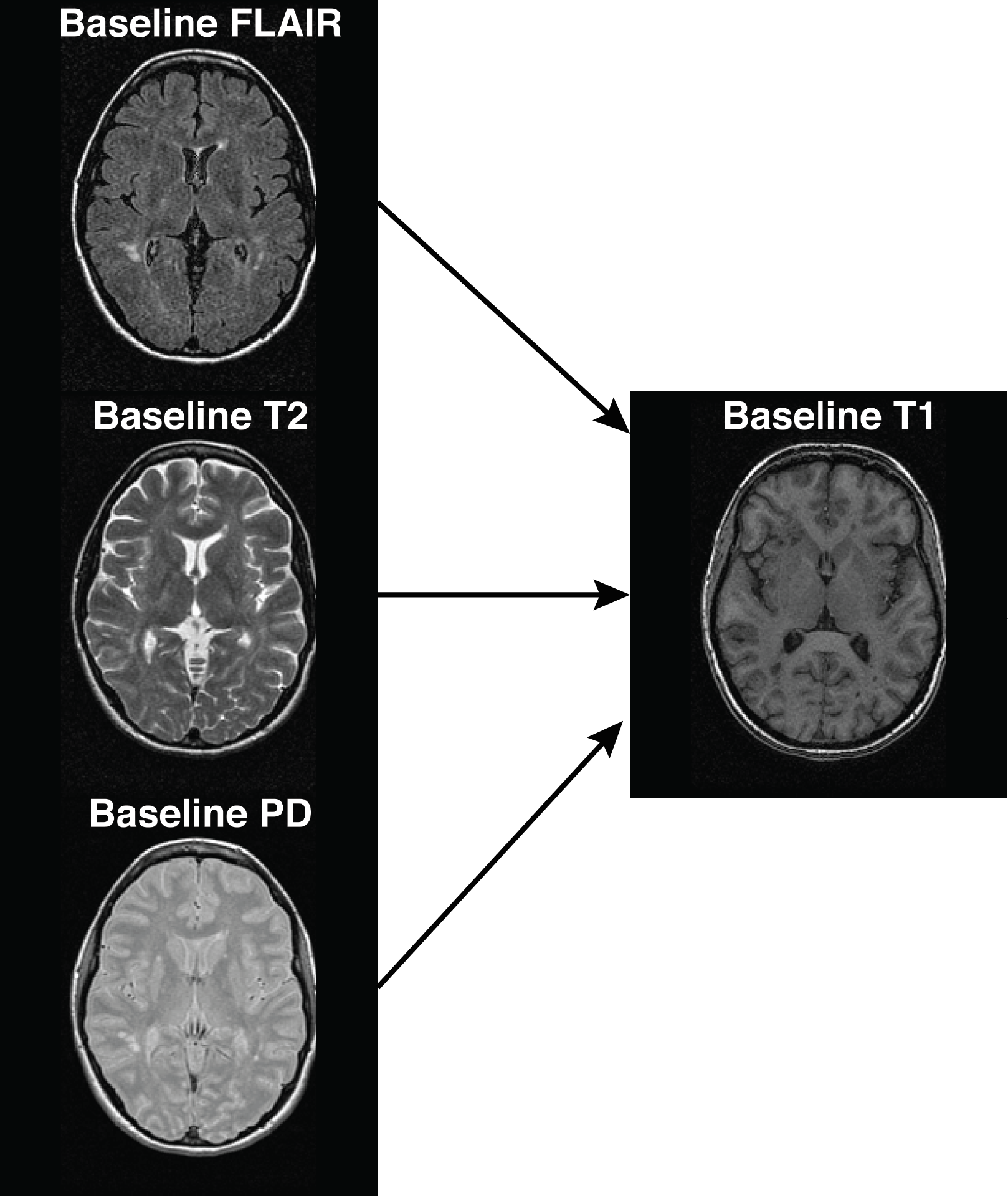 ## Registration within a visit
The function `within_visit_registration` arguments take in:
- `fixed` image - the image to be registered to
- `moving` images - images to register to the `fixed`
- `typeofTransform` - transformation of moving to fixed image (Rigid/Affine)
- `interpolator` - how are voxels averaged in `fixed` space
- `correct` - should correction be done, if so, specify `correction = "N4"`
and outputs a list of transformations (`fwdtransforms`) and output filenames (`outfile`)
## Lists: `lapply`
With `list`s and vectors, there are `apply` functions. These apply a function to every element of the list. In this course, we will use:
- `lapply` - apply function and return the elements as a `list`
- `lapply(LIST, FUNCTION, OTHER_ARGUMENTS_TO_FUNCTION)`
- `sapply` - apply function and return a "simplified" version
- if all elements returned are a one-element vector, return a `vector`
- if element 1 is a vector of length 3 and element 2 is a vector of length 1, return a `list`
## Register to the T1 image
```{r, eval = FALSE}
res = within_visit_registration(
fixed = files["T1"], moving = files[c("T2", "FLAIR")],
correct = TRUE, correction = "N4",
typeofTransform = "Rigid", interpolator = "Linear")
output_imgs = lapply(res, function(x) x$outfile)
out = c(T1 = list(t1), output_imgs)
```
```{r registration, eval = TRUE, echo = FALSE}
mods = c("T2", "FLAIR")
outfiles = file.path("..", "output", basename(files[mods]))
names(outfiles) = mods
if (!all(file.exists(outfiles))) {
res = within_visit_registration(
fixed = files["T1"],
moving = files[c("T2", "FLAIR")],
correct = TRUE, correction = "N4",
typeofTransform = "Rigid",
interpolator = "Linear"
)
output_imgs = lapply(res, function(x) x$outfile)
names(output_imgs) = mods
mapply(writenii, output_imgs, outfiles)
} else {
output_imgs = check_nifti(outfiles)
}
xout = c(T1 = list(t1), output_imgs)
mask = xout$T1 > quantile( xout$T1[ xout$T1 > 0], probs = 0.25)
dd = dropEmptyImageDimensions(mask,
other.imgs = xout)
xout = dd$other.imgs
out = lapply(xout, zscore_img, mask = dd$outimg)
out = lapply(out, window_img, window = c(-3, 3))
```
```{r reg_plot_ortho2_show, eval = FALSE}
double_ortho(out$T1, out$T2 )
```
```{r reg_plot_ortho2_run, echo = FALSE}
double_ortho(robust_window(xout$T1), robust_window(xout$T2 ))
```
## Checking registration: new visualization!
The `multi_overlay` function takes in a list of images and then plots one slice across modalities (assumes center slice):
```{r multi_overlay, echo = TRUE}
multi_overlay(out)
```
## Coregistration within a visit results
- Overall, there seems to be good overlap after registration
- It is usually beneficial to do inhomogeneity correction before registration.
- just set `correct = TRUE` or pass in the bias-corrected images
**Applying a Brain mask to all registered images**
Now that the images are in the same space as the T1, if we skull-strip the T1 image (we did with MALF), we can apply this mask to those images to extract brain tissues using the `neurobase::mask_img` command:
```{r bet_t1_show, echo = TRUE, eval = FALSE}
mask = readnii("../output/training01_01_t1_mask.nii.gz") # MALF mask
masked_imgs = lapply(out, mask_img, mask)
```
```{r bet_t1, echo = FALSE}
mask = readnii("../output/training01_01_t1_mask.nii.gz")
sub_mask = applyEmptyImageDimensions(mask, inds = dd$inds)
masked_imgs = lapply(xout, mask_img, sub_mask)
```
## Masked FLAIR Result
```{r mimgs_2}
ortho2(masked_imgs$FLAIR)
```
## Masked T2 Result
```{r mimgs_T2}
ortho2(masked_imgs$T2)
```
## Overview of functions
- Registration within a subject can be done in R
- `registration` wraps around the reading/writing of images and applying transformations (uses `ANTsR` functions)
- `double_ortho`, `ortho2`, and `multi_overlay` can provide some basic visual checks to assess registration quality
- `within_visit_registration` is a registration wrapper function
- `preprocess_mri_within` - wraps multiple steps above (inhomo, reg, extract)
- Once images are registered in the same space, operations can be applied to all the images, such as:
- Masking with a brain mask
- Transforming images to new spaces with one modality
## Co-registration overview
- Co-registration requires a few degrees of freedom (usually 6)
- sequences from the same individual/brain are more alike than images from different subjects
- Example analyses that do not require a reference template
- Identify location-specific longitudinal changes within an individual
- Tissue class or structural segmentation
- Analysis of indvidual-subject change in intensities
## Population template registration
We have only done registration within a subject, but many times you want to perform a population-level analysis. This requries registration to a **template**:
- The `registration` can be done for this as well, just the `template.file` is now the template image and `filename` is the subject image.
- other files (in the same space) can be transformed using the `other.files` and `other.outfiles` arguments. Or:
- `ants_apply_transforms` can be used to apply this transformations to the other files
## Website
http://johnmuschelli.com/imaging_in_r
## Registration within a visit
The function `within_visit_registration` arguments take in:
- `fixed` image - the image to be registered to
- `moving` images - images to register to the `fixed`
- `typeofTransform` - transformation of moving to fixed image (Rigid/Affine)
- `interpolator` - how are voxels averaged in `fixed` space
- `correct` - should correction be done, if so, specify `correction = "N4"`
and outputs a list of transformations (`fwdtransforms`) and output filenames (`outfile`)
## Lists: `lapply`
With `list`s and vectors, there are `apply` functions. These apply a function to every element of the list. In this course, we will use:
- `lapply` - apply function and return the elements as a `list`
- `lapply(LIST, FUNCTION, OTHER_ARGUMENTS_TO_FUNCTION)`
- `sapply` - apply function and return a "simplified" version
- if all elements returned are a one-element vector, return a `vector`
- if element 1 is a vector of length 3 and element 2 is a vector of length 1, return a `list`
## Register to the T1 image
```{r, eval = FALSE}
res = within_visit_registration(
fixed = files["T1"], moving = files[c("T2", "FLAIR")],
correct = TRUE, correction = "N4",
typeofTransform = "Rigid", interpolator = "Linear")
output_imgs = lapply(res, function(x) x$outfile)
out = c(T1 = list(t1), output_imgs)
```
```{r registration, eval = TRUE, echo = FALSE}
mods = c("T2", "FLAIR")
outfiles = file.path("..", "output", basename(files[mods]))
names(outfiles) = mods
if (!all(file.exists(outfiles))) {
res = within_visit_registration(
fixed = files["T1"],
moving = files[c("T2", "FLAIR")],
correct = TRUE, correction = "N4",
typeofTransform = "Rigid",
interpolator = "Linear"
)
output_imgs = lapply(res, function(x) x$outfile)
names(output_imgs) = mods
mapply(writenii, output_imgs, outfiles)
} else {
output_imgs = check_nifti(outfiles)
}
xout = c(T1 = list(t1), output_imgs)
mask = xout$T1 > quantile( xout$T1[ xout$T1 > 0], probs = 0.25)
dd = dropEmptyImageDimensions(mask,
other.imgs = xout)
xout = dd$other.imgs
out = lapply(xout, zscore_img, mask = dd$outimg)
out = lapply(out, window_img, window = c(-3, 3))
```
```{r reg_plot_ortho2_show, eval = FALSE}
double_ortho(out$T1, out$T2 )
```
```{r reg_plot_ortho2_run, echo = FALSE}
double_ortho(robust_window(xout$T1), robust_window(xout$T2 ))
```
## Checking registration: new visualization!
The `multi_overlay` function takes in a list of images and then plots one slice across modalities (assumes center slice):
```{r multi_overlay, echo = TRUE}
multi_overlay(out)
```
## Coregistration within a visit results
- Overall, there seems to be good overlap after registration
- It is usually beneficial to do inhomogeneity correction before registration.
- just set `correct = TRUE` or pass in the bias-corrected images
**Applying a Brain mask to all registered images**
Now that the images are in the same space as the T1, if we skull-strip the T1 image (we did with MALF), we can apply this mask to those images to extract brain tissues using the `neurobase::mask_img` command:
```{r bet_t1_show, echo = TRUE, eval = FALSE}
mask = readnii("../output/training01_01_t1_mask.nii.gz") # MALF mask
masked_imgs = lapply(out, mask_img, mask)
```
```{r bet_t1, echo = FALSE}
mask = readnii("../output/training01_01_t1_mask.nii.gz")
sub_mask = applyEmptyImageDimensions(mask, inds = dd$inds)
masked_imgs = lapply(xout, mask_img, sub_mask)
```
## Masked FLAIR Result
```{r mimgs_2}
ortho2(masked_imgs$FLAIR)
```
## Masked T2 Result
```{r mimgs_T2}
ortho2(masked_imgs$T2)
```
## Overview of functions
- Registration within a subject can be done in R
- `registration` wraps around the reading/writing of images and applying transformations (uses `ANTsR` functions)
- `double_ortho`, `ortho2`, and `multi_overlay` can provide some basic visual checks to assess registration quality
- `within_visit_registration` is a registration wrapper function
- `preprocess_mri_within` - wraps multiple steps above (inhomo, reg, extract)
- Once images are registered in the same space, operations can be applied to all the images, such as:
- Masking with a brain mask
- Transforming images to new spaces with one modality
## Co-registration overview
- Co-registration requires a few degrees of freedom (usually 6)
- sequences from the same individual/brain are more alike than images from different subjects
- Example analyses that do not require a reference template
- Identify location-specific longitudinal changes within an individual
- Tissue class or structural segmentation
- Analysis of indvidual-subject change in intensities
## Population template registration
We have only done registration within a subject, but many times you want to perform a population-level analysis. This requries registration to a **template**:
- The `registration` can be done for this as well, just the `template.file` is now the template image and `filename` is the subject image.
- other files (in the same space) can be transformed using the `other.files` and `other.outfiles` arguments. Or:
- `ants_apply_transforms` can be used to apply this transformations to the other files
## Website
http://johnmuschelli.com/imaging_in_r
 ## Types of Registration
- Rigid-body registration (linear) - 6 degrees of freedom (dof)
## Types of Registration
- Rigid-body registration (linear) - 6 degrees of freedom (dof)

 ## Types of Registration
- Rigid-body registration (linear) - 6 degrees of freedom (dof)
- Co-registration (within the same person)
- **Cross-sectional between-sequences**
- **Longitudinal between-sequences**
- Longitudinal within-sequence
## Types of Registration
- Rigid-body registration (linear) - 6 degrees of freedom (dof)
- Co-registration (within the same person)
- **Cross-sectional between-sequences**
- **Longitudinal between-sequences**
- Longitudinal within-sequence
 ## Types of Registration
- Rigid-body registration (linear) - 6 degrees of freedom (dof)
- Co-registration (within the same person)
- Cross-sectional between-sequences**
- Longitudinal between-sequences
- **Longitudinal within-sequence**
## Types of Registration
- Rigid-body registration (linear) - 6 degrees of freedom (dof)
- Co-registration (within the same person)
- Cross-sectional between-sequences**
- Longitudinal between-sequences
- **Longitudinal within-sequence**
 ## Types of Registration
- Affine registration – 12 dof (scaling)
- Non-linear (> 12 dof): sually requires a prior affine registration
- Across-subject registration
- Registration to a template: There are many different templates
## Types of Registration
- Affine registration – 12 dof (scaling)
- Non-linear (> 12 dof): sually requires a prior affine registration
- Across-subject registration
- Registration to a template: There are many different templates
 ## Rigid Registration: The Math
For a voxel $v$, the rigid transformation can be written as:
$$T_{\rm rigid}(v) = Rv + t$$
where $R =$
\small
$$\left[\begin{array}{ccc} \cos\beta\cos\gamma& \cos\alpha\sin\gamma + \sin\alpha\sin\beta\cos\gamma & \sin\alpha\sin\gamma - \cos\alpha\sin\beta\cos\gamma \\
-\cos\beta\sin\gamma & \cos\alpha\cos\gamma - \sin\alpha\sin\beta\sin\gamma & \sin\alpha\cos\gamma + \cos\alpha\sin\beta\sin\gamma \\
\sin\beta & -\sin\alpha\cos\beta & \cos\alpha\cos\beta \end{array}\right]$$
\normalsize
- 6 degrees of freedom
- $3$ associated with the translation vector: $t=(t_x, t_y, t_z)$
- $3$ associated with the rotation parameters: $\theta=(\alpha, \beta,\gamma)$.
## Aside into Lists
- Initialize an empty list and add two elements to it
```{r list}
l = list()
l[[1]] = c(1, 2, 4, 5)
l[[2]] = matrix(1:10, nrow = 2)
print(l)
```
- Subsetting uses double brackets:
```{r listSub}
print(l[[1]])
```
## Subsetting by name
If a `vector` has names, you can also put the
- Initialize an empty list and add two elements to it
```{r named_vec}
x = c(one = 1, three = 14, two = 5)
print(x)
x[c("three")]
```
If a `list` has names, you can subset with the `$`
- Subsetting with double brackets:
```{r named_lst}
names(l) = c("V", "m"); l[["V"]]
```
## Subsetting by name
If a `list` has names, you can also subset with the `$`
```{r named_lst_v}
l$V
```
## Reading in more than the T1: `ms.lesion` package
- `get_image_filenames_list_by_subject` - returns a list, each element is a subject, and each subject has a vector of filenames, named for each modality
```{r t1, message=FALSE}
library(ms.lesion)
all_files = get_image_filenames_list_by_subject()
class(all_files); names(all_files)
files = all_files$training01; class(files); names(files)
t1_fname = files["T1"]
t1 = readnii(t1_fname)
rt1 = robust_window(t1, probs = c(0, 0.975))
```
## Image Registration
The `registration` function from `extrantsr` can register 2 images. The main arguments are:
- `filename` - either `nifti` object or filename of image to be registered (moving)
- `template.file` - either `nifti` object or filename of target image (fixed)
- `typeofTransform` - transformation of moving to fixed image (Rigid/Affine/SyN)
- `interpolator` - how are voxels averaged in `fixed` space (Linear/NearestNeighbor/LanczosWindowedSinc)
It can also perform bias correction if `correct = TRUE`.
## Image registration
For example, if we wanted to register the FLAIR to the T1 image, we would run:
```{r, eval = TRUE, cache = TRUE, message=FALSE}
library(extrantsr)
reg = registration(
filename = files["FLAIR"], template.file = files["T1"],
typeofTransform = "Rigid", interpolator = "Linear", verbose = FALSE)
names(reg)
```
The output in `reg` would contain the transformed image and paths to the estimated transformations.
## FLAIR comparison
```{r plot_reg, eval = FALSE, cache = FALSE, message=FALSE}
double_ortho(rt1, reg$outfile)
```
```{r plot_reg_show, echo = FALSE, eval = TRUE, cache = TRUE, message=FALSE}
reg$outfile = robust_window(reg$outfile)
double_ortho(rt1, reg$outfile)
```
## Wrapper function to perform preprocessing
We would like to perform registration within a visit for all modalities. The `extrantsr` function `within_visit_registration` will do the following steps:
1. Inhomogeneity correction (N3 or N4): could also use images from previous lecture
2. Registration to a fixed image (T1)
## Rigid Registration: The Math
For a voxel $v$, the rigid transformation can be written as:
$$T_{\rm rigid}(v) = Rv + t$$
where $R =$
\small
$$\left[\begin{array}{ccc} \cos\beta\cos\gamma& \cos\alpha\sin\gamma + \sin\alpha\sin\beta\cos\gamma & \sin\alpha\sin\gamma - \cos\alpha\sin\beta\cos\gamma \\
-\cos\beta\sin\gamma & \cos\alpha\cos\gamma - \sin\alpha\sin\beta\sin\gamma & \sin\alpha\cos\gamma + \cos\alpha\sin\beta\sin\gamma \\
\sin\beta & -\sin\alpha\cos\beta & \cos\alpha\cos\beta \end{array}\right]$$
\normalsize
- 6 degrees of freedom
- $3$ associated with the translation vector: $t=(t_x, t_y, t_z)$
- $3$ associated with the rotation parameters: $\theta=(\alpha, \beta,\gamma)$.
## Aside into Lists
- Initialize an empty list and add two elements to it
```{r list}
l = list()
l[[1]] = c(1, 2, 4, 5)
l[[2]] = matrix(1:10, nrow = 2)
print(l)
```
- Subsetting uses double brackets:
```{r listSub}
print(l[[1]])
```
## Subsetting by name
If a `vector` has names, you can also put the
- Initialize an empty list and add two elements to it
```{r named_vec}
x = c(one = 1, three = 14, two = 5)
print(x)
x[c("three")]
```
If a `list` has names, you can subset with the `$`
- Subsetting with double brackets:
```{r named_lst}
names(l) = c("V", "m"); l[["V"]]
```
## Subsetting by name
If a `list` has names, you can also subset with the `$`
```{r named_lst_v}
l$V
```
## Reading in more than the T1: `ms.lesion` package
- `get_image_filenames_list_by_subject` - returns a list, each element is a subject, and each subject has a vector of filenames, named for each modality
```{r t1, message=FALSE}
library(ms.lesion)
all_files = get_image_filenames_list_by_subject()
class(all_files); names(all_files)
files = all_files$training01; class(files); names(files)
t1_fname = files["T1"]
t1 = readnii(t1_fname)
rt1 = robust_window(t1, probs = c(0, 0.975))
```
## Image Registration
The `registration` function from `extrantsr` can register 2 images. The main arguments are:
- `filename` - either `nifti` object or filename of image to be registered (moving)
- `template.file` - either `nifti` object or filename of target image (fixed)
- `typeofTransform` - transformation of moving to fixed image (Rigid/Affine/SyN)
- `interpolator` - how are voxels averaged in `fixed` space (Linear/NearestNeighbor/LanczosWindowedSinc)
It can also perform bias correction if `correct = TRUE`.
## Image registration
For example, if we wanted to register the FLAIR to the T1 image, we would run:
```{r, eval = TRUE, cache = TRUE, message=FALSE}
library(extrantsr)
reg = registration(
filename = files["FLAIR"], template.file = files["T1"],
typeofTransform = "Rigid", interpolator = "Linear", verbose = FALSE)
names(reg)
```
The output in `reg` would contain the transformed image and paths to the estimated transformations.
## FLAIR comparison
```{r plot_reg, eval = FALSE, cache = FALSE, message=FALSE}
double_ortho(rt1, reg$outfile)
```
```{r plot_reg_show, echo = FALSE, eval = TRUE, cache = TRUE, message=FALSE}
reg$outfile = robust_window(reg$outfile)
double_ortho(rt1, reg$outfile)
```
## Wrapper function to perform preprocessing
We would like to perform registration within a visit for all modalities. The `extrantsr` function `within_visit_registration` will do the following steps:
1. Inhomogeneity correction (N3 or N4): could also use images from previous lecture
2. Registration to a fixed image (T1)
 ## Registration within a visit
The function `within_visit_registration` arguments take in:
- `fixed` image - the image to be registered to
- `moving` images - images to register to the `fixed`
- `typeofTransform` - transformation of moving to fixed image (Rigid/Affine)
- `interpolator` - how are voxels averaged in `fixed` space
- `correct` - should correction be done, if so, specify `correction = "N4"`
and outputs a list of transformations (`fwdtransforms`) and output filenames (`outfile`)
## Lists: `lapply`
With `list`s and vectors, there are `apply` functions. These apply a function to every element of the list. In this course, we will use:
- `lapply` - apply function and return the elements as a `list`
- `lapply(LIST, FUNCTION, OTHER_ARGUMENTS_TO_FUNCTION)`
- `sapply` - apply function and return a "simplified" version
- if all elements returned are a one-element vector, return a `vector`
- if element 1 is a vector of length 3 and element 2 is a vector of length 1, return a `list`
## Register to the T1 image
```{r, eval = FALSE}
res = within_visit_registration(
fixed = files["T1"], moving = files[c("T2", "FLAIR")],
correct = TRUE, correction = "N4",
typeofTransform = "Rigid", interpolator = "Linear")
output_imgs = lapply(res, function(x) x$outfile)
out = c(T1 = list(t1), output_imgs)
```
```{r registration, eval = TRUE, echo = FALSE}
mods = c("T2", "FLAIR")
outfiles = file.path("..", "output", basename(files[mods]))
names(outfiles) = mods
if (!all(file.exists(outfiles))) {
res = within_visit_registration(
fixed = files["T1"],
moving = files[c("T2", "FLAIR")],
correct = TRUE, correction = "N4",
typeofTransform = "Rigid",
interpolator = "Linear"
)
output_imgs = lapply(res, function(x) x$outfile)
names(output_imgs) = mods
mapply(writenii, output_imgs, outfiles)
} else {
output_imgs = check_nifti(outfiles)
}
xout = c(T1 = list(t1), output_imgs)
mask = xout$T1 > quantile( xout$T1[ xout$T1 > 0], probs = 0.25)
dd = dropEmptyImageDimensions(mask,
other.imgs = xout)
xout = dd$other.imgs
out = lapply(xout, zscore_img, mask = dd$outimg)
out = lapply(out, window_img, window = c(-3, 3))
```
```{r reg_plot_ortho2_show, eval = FALSE}
double_ortho(out$T1, out$T2 )
```
```{r reg_plot_ortho2_run, echo = FALSE}
double_ortho(robust_window(xout$T1), robust_window(xout$T2 ))
```
## Checking registration: new visualization!
The `multi_overlay` function takes in a list of images and then plots one slice across modalities (assumes center slice):
```{r multi_overlay, echo = TRUE}
multi_overlay(out)
```
## Coregistration within a visit results
- Overall, there seems to be good overlap after registration
- It is usually beneficial to do inhomogeneity correction before registration.
- just set `correct = TRUE` or pass in the bias-corrected images
**Applying a Brain mask to all registered images**
Now that the images are in the same space as the T1, if we skull-strip the T1 image (we did with MALF), we can apply this mask to those images to extract brain tissues using the `neurobase::mask_img` command:
```{r bet_t1_show, echo = TRUE, eval = FALSE}
mask = readnii("../output/training01_01_t1_mask.nii.gz") # MALF mask
masked_imgs = lapply(out, mask_img, mask)
```
```{r bet_t1, echo = FALSE}
mask = readnii("../output/training01_01_t1_mask.nii.gz")
sub_mask = applyEmptyImageDimensions(mask, inds = dd$inds)
masked_imgs = lapply(xout, mask_img, sub_mask)
```
## Masked FLAIR Result
```{r mimgs_2}
ortho2(masked_imgs$FLAIR)
```
## Masked T2 Result
```{r mimgs_T2}
ortho2(masked_imgs$T2)
```
## Overview of functions
- Registration within a subject can be done in R
- `registration` wraps around the reading/writing of images and applying transformations (uses `ANTsR` functions)
- `double_ortho`, `ortho2`, and `multi_overlay` can provide some basic visual checks to assess registration quality
- `within_visit_registration` is a registration wrapper function
- `preprocess_mri_within` - wraps multiple steps above (inhomo, reg, extract)
- Once images are registered in the same space, operations can be applied to all the images, such as:
- Masking with a brain mask
- Transforming images to new spaces with one modality
## Co-registration overview
- Co-registration requires a few degrees of freedom (usually 6)
- sequences from the same individual/brain are more alike than images from different subjects
- Example analyses that do not require a reference template
- Identify location-specific longitudinal changes within an individual
- Tissue class or structural segmentation
- Analysis of indvidual-subject change in intensities
## Population template registration
We have only done registration within a subject, but many times you want to perform a population-level analysis. This requries registration to a **template**:
- The `registration` can be done for this as well, just the `template.file` is now the template image and `filename` is the subject image.
- other files (in the same space) can be transformed using the `other.files` and `other.outfiles` arguments. Or:
- `ants_apply_transforms` can be used to apply this transformations to the other files
## Website
http://johnmuschelli.com/imaging_in_r
## Registration within a visit
The function `within_visit_registration` arguments take in:
- `fixed` image - the image to be registered to
- `moving` images - images to register to the `fixed`
- `typeofTransform` - transformation of moving to fixed image (Rigid/Affine)
- `interpolator` - how are voxels averaged in `fixed` space
- `correct` - should correction be done, if so, specify `correction = "N4"`
and outputs a list of transformations (`fwdtransforms`) and output filenames (`outfile`)
## Lists: `lapply`
With `list`s and vectors, there are `apply` functions. These apply a function to every element of the list. In this course, we will use:
- `lapply` - apply function and return the elements as a `list`
- `lapply(LIST, FUNCTION, OTHER_ARGUMENTS_TO_FUNCTION)`
- `sapply` - apply function and return a "simplified" version
- if all elements returned are a one-element vector, return a `vector`
- if element 1 is a vector of length 3 and element 2 is a vector of length 1, return a `list`
## Register to the T1 image
```{r, eval = FALSE}
res = within_visit_registration(
fixed = files["T1"], moving = files[c("T2", "FLAIR")],
correct = TRUE, correction = "N4",
typeofTransform = "Rigid", interpolator = "Linear")
output_imgs = lapply(res, function(x) x$outfile)
out = c(T1 = list(t1), output_imgs)
```
```{r registration, eval = TRUE, echo = FALSE}
mods = c("T2", "FLAIR")
outfiles = file.path("..", "output", basename(files[mods]))
names(outfiles) = mods
if (!all(file.exists(outfiles))) {
res = within_visit_registration(
fixed = files["T1"],
moving = files[c("T2", "FLAIR")],
correct = TRUE, correction = "N4",
typeofTransform = "Rigid",
interpolator = "Linear"
)
output_imgs = lapply(res, function(x) x$outfile)
names(output_imgs) = mods
mapply(writenii, output_imgs, outfiles)
} else {
output_imgs = check_nifti(outfiles)
}
xout = c(T1 = list(t1), output_imgs)
mask = xout$T1 > quantile( xout$T1[ xout$T1 > 0], probs = 0.25)
dd = dropEmptyImageDimensions(mask,
other.imgs = xout)
xout = dd$other.imgs
out = lapply(xout, zscore_img, mask = dd$outimg)
out = lapply(out, window_img, window = c(-3, 3))
```
```{r reg_plot_ortho2_show, eval = FALSE}
double_ortho(out$T1, out$T2 )
```
```{r reg_plot_ortho2_run, echo = FALSE}
double_ortho(robust_window(xout$T1), robust_window(xout$T2 ))
```
## Checking registration: new visualization!
The `multi_overlay` function takes in a list of images and then plots one slice across modalities (assumes center slice):
```{r multi_overlay, echo = TRUE}
multi_overlay(out)
```
## Coregistration within a visit results
- Overall, there seems to be good overlap after registration
- It is usually beneficial to do inhomogeneity correction before registration.
- just set `correct = TRUE` or pass in the bias-corrected images
**Applying a Brain mask to all registered images**
Now that the images are in the same space as the T1, if we skull-strip the T1 image (we did with MALF), we can apply this mask to those images to extract brain tissues using the `neurobase::mask_img` command:
```{r bet_t1_show, echo = TRUE, eval = FALSE}
mask = readnii("../output/training01_01_t1_mask.nii.gz") # MALF mask
masked_imgs = lapply(out, mask_img, mask)
```
```{r bet_t1, echo = FALSE}
mask = readnii("../output/training01_01_t1_mask.nii.gz")
sub_mask = applyEmptyImageDimensions(mask, inds = dd$inds)
masked_imgs = lapply(xout, mask_img, sub_mask)
```
## Masked FLAIR Result
```{r mimgs_2}
ortho2(masked_imgs$FLAIR)
```
## Masked T2 Result
```{r mimgs_T2}
ortho2(masked_imgs$T2)
```
## Overview of functions
- Registration within a subject can be done in R
- `registration` wraps around the reading/writing of images and applying transformations (uses `ANTsR` functions)
- `double_ortho`, `ortho2`, and `multi_overlay` can provide some basic visual checks to assess registration quality
- `within_visit_registration` is a registration wrapper function
- `preprocess_mri_within` - wraps multiple steps above (inhomo, reg, extract)
- Once images are registered in the same space, operations can be applied to all the images, such as:
- Masking with a brain mask
- Transforming images to new spaces with one modality
## Co-registration overview
- Co-registration requires a few degrees of freedom (usually 6)
- sequences from the same individual/brain are more alike than images from different subjects
- Example analyses that do not require a reference template
- Identify location-specific longitudinal changes within an individual
- Tissue class or structural segmentation
- Analysis of indvidual-subject change in intensities
## Population template registration
We have only done registration within a subject, but many times you want to perform a population-level analysis. This requries registration to a **template**:
- The `registration` can be done for this as well, just the `template.file` is now the template image and `filename` is the subject image.
- other files (in the same space) can be transformed using the `other.files` and `other.outfiles` arguments. Or:
- `ants_apply_transforms` can be used to apply this transformations to the other files
## Website
http://johnmuschelli.com/imaging_in_r